Hierarchical Clustering in a Nutshell
The following image, taken from the Wikipedia article on HC, illustrates the basic notion rather well:
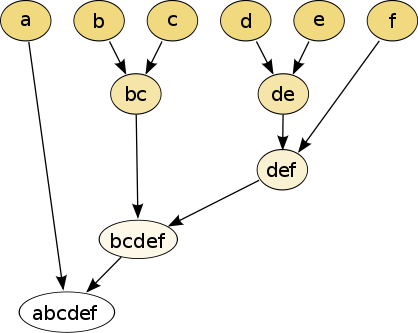
Suppose we have 6 items that we wish to cluster, and we have some metric that we can compute between subsets of those items. We get a hierarchy of clusters via the following greedy algorithm:
- Let each item be a cluster (with a single element)
- Let $S$ be the set of all clusters
- While $|S| > 1$:
- Find the closest pair of clusters $X, Y$ in $S$.
- Define a new cluster $Z := X \cup Y$
- Add $Z$ to $S$, remove $X$ and $Y$ from $S$
So the diagram above implies the following possible sequence of cluster unions (it may be slightly different):
- $\{b\} \cup \{c\}$
- $\{d\} \cup \{e\}$
- $\{d,e\} \cup \{f\}$
- $\{b,c\} \cup \{d,e,f\}$
- $\{a\} \cup \{b,c,d,e,f\}$
There are a number of obvious optimizations, for example, one does not have to recompute all distances after each creation of a new cluster. Rather - only the distances that involve the new cluster.
What is It Good For?
In most of the use cases that we employed, the initial atoms were triangular facets of a 3D mesh. One such use case is mesh segmentation, that is the process of breaking down a 3D object into meaningful sub-parts. There's a well-known paper from 2006 by Attene et al that describes such an approach. The metric chosen for distance between segments(=clusters) is how much they resemble certain primitive shapes the authors chose in advance (cylinder, cube, sphere, etc.).
As can be seen in this image taken from the paper, once one has the full hierarchy of segments, this tree can be trimmed according to the number of clusters one wishes.
 |
| source: "Hierarchical mesh segmentation based on fitting primitives" by Attene et al |
The Problem With HC
The natural approach we took was to consider this not as atoms where all pairwise distances needed to be considered, but as a graph, where only distances between neighboring vertices had to be considered. In the 3D mesh case - adjacent faces were represented by neighboring atoms in the HC tree. So now we simply put all the edges of the graph (and the respective distances) into a some implementation of a min-heap, and began the simple process of:
- Extracting the minimal edge
- Uniting the clusters it connected
- Updating the graph (and the edges in the heap) accordingly
This became an issue when the number of items stored in the heap was in the millions and tens of millions, which is very much the case when you get a high-quality 3D-scan of a room, for example. It turned out that operations on the heap became unbearably expensive, and the memory footprint was terrible, since we had to store this huge heap in the RAM.
The Solution: HHC
We realized that changes in the HC tree were almost always very local - one computes the distance between pairs of clusters, which are neighboring nodes in the graph, and sometimes unites them in a way which affects only the neighbors of the united clusters. So why not divide and conquer?
What we ended up doing is this:
- Break down the model very crudely into K parts, by doing something that is not far from just putting it on a grid and taking cubes. Choose K such that each part will have not so many facets in it, but not too little.
- Run HC on each part separately until the number of clusters in the part becomes small.
- Now unite adjacent parts until the number of clusters in each part is again not too big but not too small.
Note that this is in effect doing a Hierarchical Clustering of the parts, hence the winning name.
Also note that it effectively means you work on a number of very small heaps all the time and never on a very large one. This means heap operations are now considerably cheaper, and indeed the memory footprint went down by a factor of 5 or so, and the running time was improved dramatically, on machines with slower hard drives (since less swapping was involved).
The Price: Accuracy
As it often happens, the price of heuristics is that they mess up the algorithm's accuracy. In our case - it means that if the minimal edge happens to connect atoms that lie in different parts - you will get to it much later than you otherwise would have, but the reduction in running time and resource consumption made it worth our while.